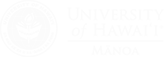The Development of New Multidentate, Non-innocent, and/or Chiral Ligands for Application in Transition Metal-based Asymmetric Catalysis, Small Molecule Activation, and Alternative Energy Processes — Cain
My research interests include development of new multidentate, non-innocent, and/or chiral ligands for application in transition metal-based asymmetric catalysis, small molecule activation, and alternative energy processes. Without giving up too much information, we plan to target a largely unexplored class of C1-Symmetric, P-Stereogenic ligands, rationally designed multidentate ligands for dinitrogen reduction, and phosphaalkene-based ligands as redox active / non-innocent platforms for interesting chemical transformations. If these projects interest you, please do not hesitate to contact me or stop by my office.
For more information, please visit our group website, Synthesis in Paradise.
— Fuller
Folding Events in Proteins — Haglund
Ellinor’s research is focused on the folding and function of proteins, utilizing both computational and experimental techniques to understand the molecular details of how proteins fold into biologically active molecules. Through her postdoctoral study she discovered a new class of “knot-like” motifs called pierced lasso topologies (PLTs). Interestingly, these threaded topologies exist in all kingdoms of life, with 14 different biological functions, and populate about 18% of known protein structures containing a disulphide bond. Ellinor’s research focuses on three strategically selected model systems (i) the pleiotropic hormone leptin and the leptin receptor, (ii) the oxidoreductase superoxide dismutase, and (iii) chemokine ligands and their cognate receptors. The long-term goal is to understand the biological relevance of its PLT and their role in human health and disease. Ellinor is inspired by how nature works and utilize my multidisciplinary training to answer questions at the interface of chemistry, biology, and physics.
Organoelement Chemistry and Synthesis of Hypervalent Organobismuth Complexes — Hyvl
My research interests include organoelement chemistry with emphasis on exploration of new synthetic transformations, reactivity and applications. The research in my group will focus on synthesis of hypervalent organobismuth complexes supported with specially-designed ligand scaffolds for their utilization in transition-metal catalyzed cross-couplings, small molecule activation and generation of metallodrugs.
For more information about the Hyvl Research Group, please visit our website: https://sites.google.com/a/hawaii.edu/hyvl/
The S-Adenosylmethionine-Dependent Radical Enzymes — Jarrett
Research Interests
The S-Adenosylmethionine-Dependent Radical Enzymes
Organic radicals are used by a number of enzymes to catalyze biochemical transformations with high-energy barriers that would be difficult to accomplish through non-radical heterolytic chemistry. Well known examples include the reduction of an alcohol to an alkane catalyzed by ribonucleotide reductase or carbon chain rearrangements catalyzed by methylmalonyl CoA mutase or glutamate mutase. Organic radicals can be generated in enzymes through only three general schemes: metal-activated oxygen chemistry, adenosylcobalamin chemistry, or reduction of the sulfonium of S-adenosylmethionine (AdoMet).
Enzymes that catalyze the reduction of AdoMet all contain a [4Fe-4S] cluster that interacts covalently with the methionyl portion of AdoMet. This interaction likely plays an important role in promoting electron transfer into the relatively low-potential sulfonium. We are interested in understanding how the electronic structure of the FeS cluster and the sulfonium are perturbed by this interaction, and whether electron transfer and sulfonium reduction requires a structurally specific interaction or simply close proximity.
The AdoMet Radical Enzyme Biotin Synthase
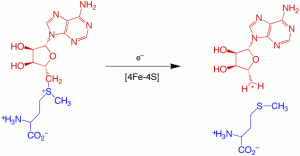
Biotin (vitamin H) is synthesized in microbes and plants through a conserved biosynthetic pathway that generates dethiobiotin from L-alanine, pimeloyl CoA, CO2, and NH3 (from AdoMet). The final step in the pathway requires the substitution of sulfur for hydrogen at the C6 and C9 positions to create the thiophane ring. The enzyme that catalyzes this step in the reaction contains iron-sulfur clusters and is an AdoMet-dependent radical enzyme. The overall reaction catalyzed is:
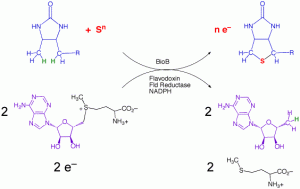
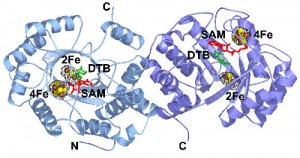
In collaboration with Dr. Cathy Drennan (MIT), we have determined the structure of the enzyme from E. coli with substrates bound. On the basis of this structure, and related mechanistic studies, we have proposed that the biotin thiophane sulfur is derived from a [2Fe-2S] cluster bound within the core of the enzyme. Current studies are focused on obtaining spectroscopic evidence for a dethiobiotin-FeS cluster covalent intermediate and understanding the role of conserved protein residues in guiding this transformation.
In Vivo Iron-Sulfur Cluster Assembly
Although simple iron-sulfur clusters are spontaneously assembled in solution, virtually all known organisms contain specific proteins that function to guide the controlled assembly of [2Fe-2S] and [4Fe 4S] clusters. One of the most conserved systems for iron-sulfur cluster assembly, the ISC system, is contained within a single genetic operon in E. coli that codes for 7 proteins: a pyridoxal phosphate-dependent cysteine desulfurase that generates S0 as persulfide sulfur destined for cluster assembly (IscS), two possible scaffold proteins that can bind either [2Fe-2S], [4Fe 4S], or Fe2+/3+ under various experimental conditions (IscU and IscA), a ferredoxin for redox control (Fdx), and an ATP-dependent chaperone/co-chaperone pair that may assist in folding/unfolding or cluster transfer (HscA and HscB).
We have found that the chaperone HscA and the scaffold IscU will form a complex with several apoproteins that are targets for FeS cluster transfer. Our current studies are focused on cluster assembly in biotin synthase, which is stably folded in the apoprotein state, and where correct cluster assembly can be distinguished from indiscriminant assembly by the ability of the resulting protein to form biotin. Current studies focus on using spectroscopic methods to characterize the state of the FeS cluster prior to and during the cluster transfer process.
Potential projects:
- Model studies of sulfonium reduction chemistry.
- Re-engineering biotin synthase to contain a spectroscopic probe to follow AdoMet cleavage and radical generation.
- Mutagenesis of conserved protein residues in biotin synthase to probe the role of the enzyme in guiding the radical chemistry.
- Use of deuterated dethiobiotin to determine isotope effects for the cleavage of C-H bonds by biotin synthase, to better understand the transition state structure.
- Spectroscopic and electrochemical studies of cluster assembly and transfer within protein members of the ISC system.
- Mutagenesis of conserved protein residues in ISC proteins to probe the role of the proteins in controlling cluster assembly and targeting to specific apoproteins.
Development of Catalytically Enhanced Complex Aluminum Hydrides as Vehicular Hydrogen Storage Materials — Jensen
Reaction Dynamics and Materials in Extreme Environments — Kaiser
Applications and Methodology Development of High-resolution Solid-state Nuclear Magnetic Resonance (NMR) for the Study of Novel Systems — Kumashiro
Her research program is centered about the applications and methodology development of high-resolution solid-state nuclear magnetic resonance (NMR) for the study of novel systems, such as connective tissue proteins. Extramural support for Dr. Kumashiro’s research program includes a CAREER award from the National Science Foundation.
Current research projects focus on elastin,a vertebrate protein with remarkable biomechanical properties. The elastic properties of a number of physiological structures, such as blood vessels and skin, are believed to originate from elastin, an amorphous, crosslinked protein comprised largely of small hydrophobic amino acids. The three-dimensional structures of insoluble elastin remains elusive, as crystallographic tools and even the most sophisticated solution NMR spectroscopy cannot be used to study this insoluble protein. Therefore, for many years, the true nature of elasticity in biological systems has remained controversial, as none of the existing models could be confirmed or rebutted with high-resolution structural data.
Now, with the advent of high magnetic fields and a growing number of high-resolution solid-state NMR experiments, the structural characterization of elastin is in progress in our laboratories. Solid-state NMR has proven to be a powerful tool in the characterization of molecular and bulk properties of many compounds. For instance, isotropic chemical shift and chemical shift anisotropy are relative measures of the shielding of a specific nucleus and are sensitive probes of chemical or electronic environments. More elaborate experiments are used to ascertain other structural parameters, such as internuclear distances or torsion angles, as demonstrated by others for a range of peptides and proteins. In addition, a number of experiments are used to determine dynamical features. These experiments include, e.g., relaxation measurements that are analogous to those done in the solution state. Finally, in addition to the application of now-routine solid-state NMR experiments for structural studies of proteins and polymers, we have also endeavored to develop, adapt, and apply new methods for these samples.
We have adopted an approach that includes use of a wide range of elastin and elastin peptide samples. In this manner, we anticipate that a global model for biological elasticity will emerge, rather than a set of observations that may or may not be specific for a given elastin-based motif.
One set of work focuses on characterization of the natural-abundance 13C populations in elastin isolated from tissue, in addition to elastin peptides from numerous collaborators. With regards to the latter, these ongoing collaborations include elastin peptides from synthesis as well as recombinant methods.
In addition to the natural-abundance 13C work, we have developed a unique approach to isotopic labeling of this vertebrate protein. As bacterial expression methods lack the natural mechanism for crosslinking, we have adapted a protocol using a mammalian cell line to produce insoluble elastin with 13C, 15N, and 2H-labels at key amino acids. Not only does this approach provide unambiguous structural assignments for the enriched sites, but it increases the number of NMR experiments that are now feasible for this complex system.
Our early work, which includes both the natural-abundance and isotopically-enriched preparations, highlight the unusual mobility of large segments of elastin. Furthermore, our data show strong support for models of elastin structure that involve significant structural heterogeneity (rather than regular or repeating structures). The curious reader is encouraged to consult recent publications from this lab for additional information.
Atmospheric chemistry, astrochemistry, combustion chemistry, and superfluidity — Raston
Research in the Raston Lab is focused on advancing our understanding of atmospheric chemistry, astrochemistry, combustion chemistry, and superfluidity. We use a variety of home-built spectrometers in order to do this, that span the lower end of the electromagnetic spectrum (microwave to infrared). We are currently focusing on two projects. The first involves preparing a variety of exotic chemical intermediates in both the gas phase and in helium nanodroplets. The spectroscopy we plan to perform on them should provide insight into their reactivity, in addition to rest frequencies to help with their identification in various environments (such as the interstellar medium). The second project focuses on investigating the quantum solvation of different types of molecules with either helium or hydrogen. Through the power of high-resolution spectroscopy we hope to uncover new manifestations of microscopic superfluidity in these types of systems. We’re also developing a variety of scientific instrument simulator’s that will provide students with virtual lab experiences when in-person labs are not possible.
For more information, please visit our groups website at https://rastonlab.org.
The Role of Spin-orbit Coupling and Rotational Excitation on Ion-molecule Reactions, Modeling of a Realistic Multi-component Membrane, and Realistic Multi-component Membrane Dynamics. — Sun
For more information on the Sun Research Group, please click here.
Organic Synthesis — Tius
For more information please see:
Natural Product Drug Discovery — Williams
Natural Product Drug Discovery
Despite the long history of drug discovery from natural sources, the marine environment is still relatively untapped. Covering 70% of the Earth’s surface, the oceans contain all major phyla. The resulting intense competition for space and resources drives the evolution of specific and potent chemical defenses distinct from their terrestrial counterparts.
My research interests center on the discovery and evaluation of these small molecule chemical defenses from marine sources as potential drug leads for the treatment of cancers and Alzheimer’s Disease.
In collaboration with the other members of the Natural Products Cancer Biology program, marine extracts are screened against a variety of relevant cancer targets. The active constituents are then isolated using a combination of bioassay data and repeated separations. The structures of these metabolites are then determined primarily through the use of high-field NMR spectroscopy and chemical degradation.
New Natural Product Methodologies
Modern structure determination of small molecules still remains a challenging problem. The number of structural reassignments reported each year is testimony to the myriad of potential pitfalls. While classical approaches to structure determination rely heavily on chemical degradation, most modern approaches use non-destructive techniques to provide the same connectivity information. Any structure determination encompasses three discrete assignments: planar, relative and absolute. Advances in NMR instrumentation and NMR pulse sequences have greatly simplified the assignment of the planar structure, i.e., the constitutional connectivities between the various nuclei. Conversely, relative and absolute stereochemical assignments are becoming more challenging as the isolation of submicromolar quantities of metabolites becomes increasingly more common. Consequently, my group focuses on developing new methodologies for stereochemical determinations on small scale.
More information about the Williams Research group can be found here
Organic synthesis methodologies development via merging both radical and polar reactivity — Zhang
Zuxiao’s research is focused on the drug discovery directed organic synthesis methodology development. Through his postdoctoral study he realized regio-, and enantioselective C-H functionalization via the merger of “radical chaperone” strategy and copper catalyzed radical relay. Zuxiao’s research focuses on three directions, fluorine chemistry, radical chemistry, and asymmetric catalysis. The long-term goal is to develop novel catalytic system by harness both radical and polar reactivity to realize selective functionalization of inert chemical bonds initiated multicomponent reactions, thus to provide an efficient way to access biorelevant molecules, as well as a robust tool for late-stage functionalization of complex drug molecules
For more information on the Zhang Research Group, please click here.