New technology helps researchers trace origins of codfish decline
Scientists can now see how climate change affects the copepod, a key food source for cod
University of Hawaiʻi at MānoaResearcher, Pacific Biosciences Research Center
Andrew Christie, (808) 956-5212
Associate Research Specialist, Pacific Biosciences Research Center
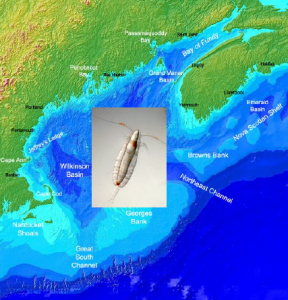
The mega-decline in cod and other fisheries across the North Atlantic Ocean threatens the livelihood of fishermen and communities in New England and Atlantic Canada. One suspect in the disappearance of cod and other groundfish is the food source for their young: a planktonic copepod crustacean, no larger than a grain of rice. Recent changes in local copepod populations have co-occurred with declines in fisheries elsewhere, such as the collapse of the cod fishery in Europe’s North Sea.
For this and other reasons, scientists at the University of Hawai‘i at Mānoa are working to understand how copepods are responding to global climate change, including to increases in water temperature, altered ocean currents, and ocean acidification.
Because of the copepod’s small size and its vast ocean habitat, it is a poor subject for conventional physiological studies. New molecular techniques have opened doors for an alternative approach. Known as transcriptomics, this technique makes a catalog of all the messages (“transcripts”) produced by the cells that control the animal’s physiology. With this tool, biologists are now able to listen in on the instructions being sent out to direct an organism’s response to its changing environment. With respect to copepods, the challenge is to identify and understand each message, in order to track down the causes of population changes.
The first transcriptome for the key North Atlantic copepod Calanus finmarchicus has now been published and made available for scientists everywhere. Appearing in a February 2014 issue of the journal PLOS ONE, it is the work of a team of scientists from the Pacific Biosciences Research Center at UH Mānoa, Ohio University and Indiana University’s National Center for Genome Analysis Support. Mount Desert Island Biological Laboratory in Salisbury Cove, Maine provided access to the biological samples. The transcriptome was sequenced by the University of Georgia Genomics Facility in Athens, Georgia.
This publication provides the first publicly accessible, large-scale molecular resource for investigating the physiological ecology of Calanus. Highlights of this study include: 1) the observation that a large percentage of genes are not in play at any particular time while a young copepod matures; 2) discovery of specific messages being sent out as individuals prepare to enter a critical dormant phase in their annual population cycle; and 3) the discovery of a number of previously unknown genes, suggesting a more complex biology than that of related animals like the fruit fly and water flea, which are used extensively for biomedical and ecotoxicologial research.
With the Calanus transcriptome in hand, scientist now have a tool to better understand how copepods adapt, and may be better able to predict when and where population changes will occur for this planktonic crustacean on which many fisheries depend.
# # #
The original article describing this study can be accessed free of charge at: http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0088589
Several presentations describing or using this new resource will be presented at the 2014 Ocean Sciences Meeting being held February 23-28 at the Hawaii Convention Center, in Honolulu, Hawaii. Funding for this project was provided by the National Science Foundation and the Cades Foundation of Honolulu, Hawaiʻi.
For more information, visit: http://www.pbrc.hawaii.edu/